Advanced Example 1: Mutli-Coil#
This notebook uses the concepts shown in the previous introductions, to build a more advanced simulation using multiple coils.
The structure should look familiar to the introductions, namely: - Define simulation - Set up digital phantom - Run simulation and show results
Import#
external modules
[1]:
import sys
sys.path.append('../../')
sys.path.append("../../tests/")
import tensorflow as tf
physical_devices = tf.config.list_physical_devices('GPU')
if physical_devices:
print("Available GPUS: \n\t", "\n\t".join([str(_) for _ in physical_devices]))
tf.config.experimental.set_visible_devices(physical_devices[0], 'GPU')
tf.config.experimental.set_memory_growth(physical_devices[0], True)
from collections import OrderedDict
from typing import Union, Iterable, Tuple
import numpy as np
import math
import matplotlib.pyplot as plt
%matplotlib inline
import cmrsim
import phantoms
Available GPUS:
PhysicalDevice(name='/physical_device:GPU:0', device_type='GPU')
Define Simulation#
This subclassed simulation does not only implement the required abstract methods, but also implements functions to determine the noise-level for a given SNR within the left ventricle of the myo-cardium. The digital phantom, used for simulation is a slice from the XCAT phantom again.
The _build method does not define a reconstruction module, as this is performed inside the function delayed_reconstruction.
[2]:
class MultiCoilSET2star(cmrsim.analytic.AnalyticSimulation):
def build(self, fov: Tuple[int, int], kspace_dims: Tuple[int, int], TE: float, TR: float, n_coils: int = 8):
n_kx, n_ky = kspace_dims
# Encoding definition
segments = [kspace_dims[1]//i for i in range(1, int(math.sqrt(kspace_dims[1]))+1) if kspace_dims[1] % i == 0][-2]
encoding_module = cmrsim.analytic.encoding.EPI(field_of_view=fov, sampling_matrix_size=(n_kx, n_ky),
read_out_duration=0.1, blip_duration=0.01,
k_space_segments=27, absolute_noise_std=0.)
# Signal module construction
spinecho_module = cmrsim.analytic.contrast.SpinEcho(echo_time=TE, repetition_time=TR)
mdims = np.array([(-fov[0]/2, fov[0]/2), (-fov[1]/2, fov[1]/2), (0., 0.1)])
coil_sens_module = cmrsim.analytic.contrast.BirdcageCoilSensitivities(map_shape=np.array((n_kx*4, n_ky*4, 1, 8)).astype(int),
map_dimensions=mdims)
t2_star_module = cmrsim.analytic.contrast.StaticT2starDecay(encoding_module.get_sampling_times())
# Forward model composition
forward_model = cmrsim.analytic.CompositeSignalModel(spinecho_module, coil_sens_module, t2_star_module)
return forward_model, encoding_module, None
def _update(self):
super()._update()
self.forward_model.static_t2star.sampling_times = self.encoding_module.get_sampling_times()
def estimate_noise_level(self, simulated_k_space: tf.Tensor, mask: Union[np.ndarray, tf.Tensor],
target_snr: np.ndarray):
""" This function is not defined in the base class but serves the porpose of accesing the modules
and its parameters that are within the scope of the simulation instance!
It will perform a channelwise (per coil) reconstruction and then estimate the noise-level needed to
yield the target SNR in the Roemer-reconstructed Multi-coil images.
"""
sampling_matrix_size = self.encoding_module.matrix_size.read_value()
coil_sens_maps = tf.transpose(self.forward_model.coil_sensitivities.coil_maps, (3, 0, 1, 2))
inverse_crime_factor = tf.cast(coil_sens_maps.shape[1], tf.int32) // sampling_matrix_size[0]
channel_wise_recon = cmrsim.reconstruction.Cartesian2D(sample_matrix_size=sampling_matrix_size)
single_coil_images = channel_wise_recon(simulated_k_space)
unstacked_channelwise_images = self.forward_model.unstack_repetitions(single_coil_images)
print(single_coil_images.shape, unstacked_channelwise_images.shape, coil_sens_maps.shape)
downsampled_coilmaps = coil_sens_maps[:, ::inverse_crime_factor, ::inverse_crime_factor, 0]
downsampled_masks = mask[::inverse_crime_factor, ::inverse_crime_factor].astype(np.float32)
print(downsampled_coilmaps.shape, downsampled_masks.shape)
noise_std_deviations = cmrsim.utils.snr.compute_noise_std(
noiseless_single_coil_images=unstacked_channelwise_images[:, 0, ...],
coil_sensitivities=downsampled_coilmaps,
target_snr=target_snr,
mask=downsampled_masks,
iteratively_refine=False)
return noise_std_deviations, single_coil_images
def delayed_reconstruction(self, simulated_k_space: tf.Tensor):
""" This method performs a Roemer reconstruction outside of the Graph """
sampling_matrix_size = self.encoding_module.matrix_size.read_value()
coil_sens_maps = tf.transpose(self.forward_model.coil_sensitivities.coil_maps, (3, 0, 1, 2))
inverse_crime_factor = tf.cast(coil_sens_maps.shape[1], tf.int32) // sampling_matrix_size[0]
channel_wise_recon = cmrsim.reconstruction.Cartesian2D(sample_matrix_size=sampling_matrix_size)(simulated_k_space)
downsampled_coilmaps = tf.transpose(coil_sens_maps[:, ::inverse_crime_factor, ::inverse_crime_factor, 0], (0, 2, 1))
recon_module = cmrsim.reconstruction.Cartesian2DMultiCoil(sample_matrix_size=sampling_matrix_size,
coil_sensitivities=downsampled_coilmaps)
return recon_module(simulated_k_space, coil_channel_axis=0), channel_wise_recon
Instantiate the simulator
[3]:
From lookup base (1, 480, 360, 1, 16)
Load the semantic label map and assign the relaxation_times
[4]:
phantom_dict, phantom_reference_map, coordinates = phantoms.define_shepplogan(fov, phantom_matrix)
phantom_dict["T2star"] = np.ones_like(phantom_dict["T1"]) * 35.
# phantom_dict["r_vectors"] = np.stack([phantom_dict["r_vectors"][..., i] for i in (1, 0, 2)], axis=-1)
for k, v in phantom_dict.items():
if k != "r_vectors":
phantom_dict[k] = v.reshape(-1)
print(phantom_reference_map.shape, [f"{k}: {v.shape}" for k, v in phantom_dict.items()])
dataset = cmrsim.datasets.AnalyticDataset(phantom_dict, filter_inputs=True, expand_dimension=True)
(360, 480) ['M0: (172800,)', 'r_vectors: (172800, 3)', 'T1: (172800,)', 'T2: (172800,)', 'T2star: (172800,)']
Run the simulation#
[5]:
%time noise_less_k_space = simulator(dataset, voxel_batch_size=2500, execute_eager=True)
Run Simulation: |XXXXXXXXXXXXXXX|92372/92372 |XXXXXXXXXXXXXXX|27/27 CPU times: total: 1min 12s
Wall time: 20.9 s
Estimate Noise levels and add noise to the k-space data
[6]:
print('Noiseless shape: ', noise_less_k_space.shape)
target_snrs = [100, 20, 5]
estimated_noise_levels, single_coil_images = simulator.estimate_noise_level(noise_less_k_space,
phantom_reference_map.T > 0,
target_snrs)
print('Single coil shape: ', single_coil_images.shape)
simulator.encoding_module.absolute_noise_std.assign(tf.constant(estimated_noise_levels))
simulator._update()
# Note: second-to-last axis in noise-less-k-space already is a "noise-axis". That why the indexing is used.
# Re-running the simulation at this point would yield the same result
noisy_k_space = simulator.encoding_module.add_noise(noise_less_k_space[..., 0, :])
print("Noisy shape: ", noisy_k_space.shape)
combined_images, noisy_single_channel_images = simulator.delayed_reconstruction(noisy_k_space)
print("Images shape: ", combined_images.shape, noisy_single_channel_images.shape)
Noiseless shape: (8, 1, 10800)
(8, 1, 90, 120) (8, 1, 90, 120) (8, 480, 360, 1)
(8, 120, 90) (120, 90)
Single coil shape: (8, 1, 90, 120)
Noisy shape: (8, 3, 10800)
Images shape: (3, 90, 120) (8, 3, 90, 120)
Plot images:#
[11]:
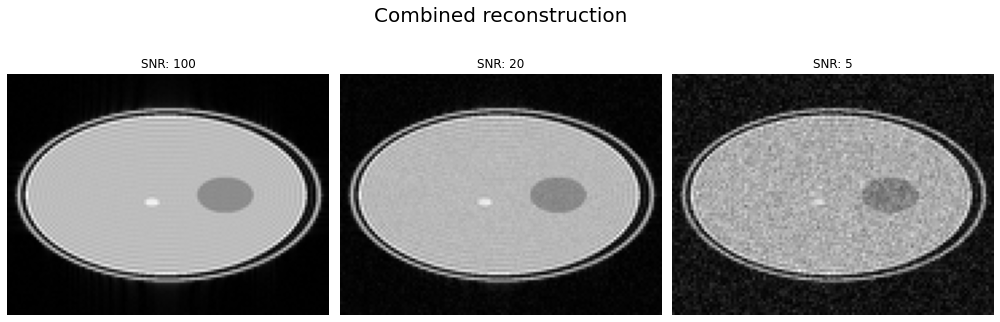
with accordion.children[0]:
f, axes = plt.subplots(1, 3, figsize=(14, 5))
magnitude = np.abs(np.squeeze(combined_images))
for snr_idx, a, img in zip(range(magnitude.shape[1]), axes, magnitude):
a.axis("off")
a.imshow(img, cmap="gray")
a.set_title(f"SNR: {target_snrs[snr_idx]}")
f.suptitle("Combined reconstruction", fontsize=20)
f.tight_layout()
[13]:
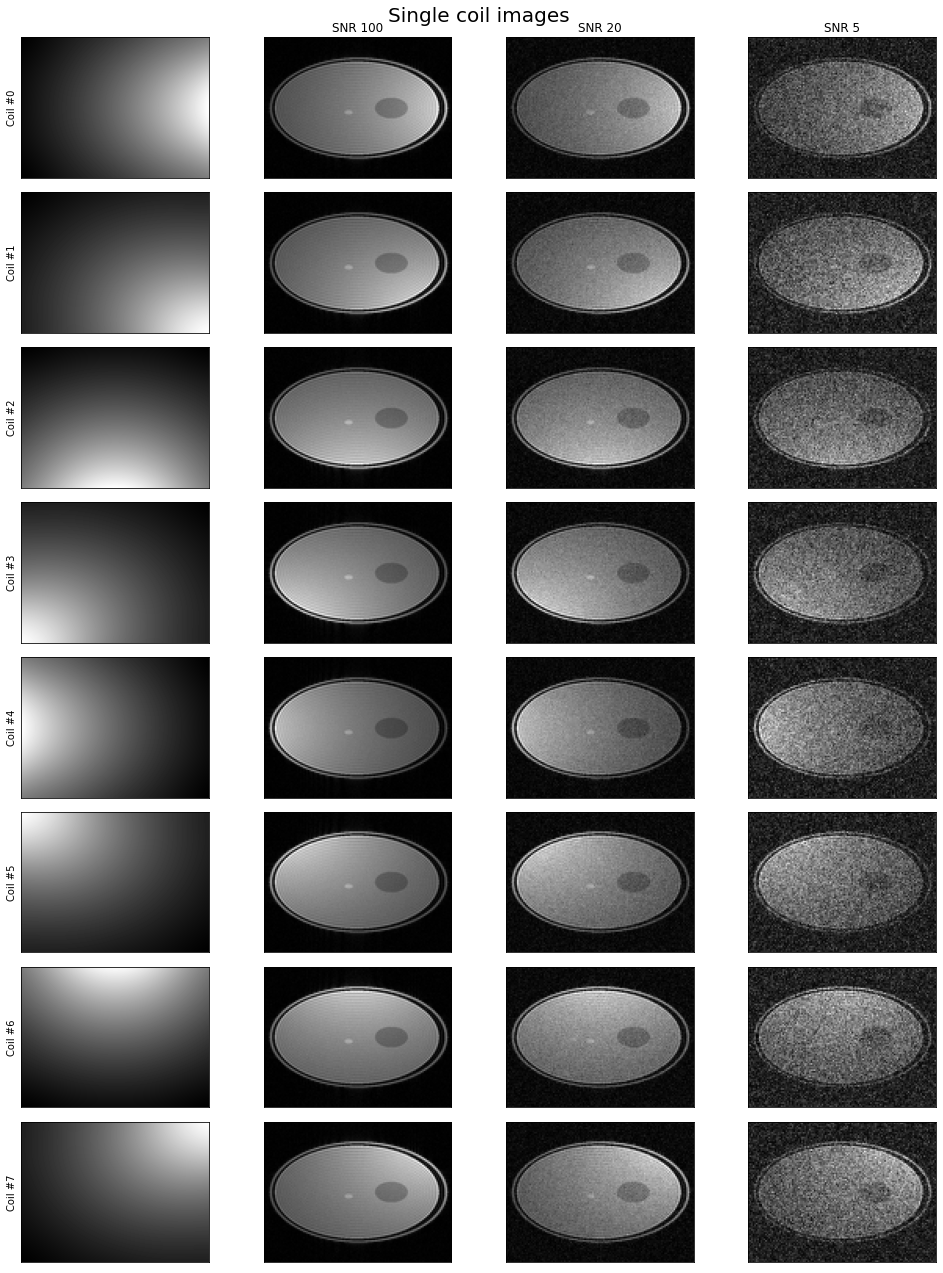
n_cols = 1 + noisy_single_channel_images.shape[1]
f, axes = plt.subplots(8, n_cols, figsize=(3.5 * n_cols, 18))
coil_magnitude = np.abs(np.squeeze(simulator.forward_model.coil_sensitivities.coil_maps))
for cidx, a, img in zip(range(8), axes[:, 0], coil_magnitude.transpose(2, 0, 1)):
a.set_yticks([]), a.set_xticks([])
a.imshow(img.T, cmap="gray")
a.set_ylabel(f"Coil #{cidx}")
magnitude = np.abs(noisy_single_channel_images)
for snr_idx in range(magnitude.shape[1]):
axes[0, snr_idx+1].set_title(f"SNR {target_snrs[snr_idx]}")
for coil_idx, a in zip(range(magnitude.shape[0]), axes[:, snr_idx+1]):
a.set_yticks([]), a.set_xticks([])
a.imshow(magnitude[coil_idx, snr_idx], cmap="gray")
f.suptitle( "Single coil images", fontsize=20), f.tight_layout()
[13]:
(Text(0.5, 0.98, 'Single coil images'), None)
[ ]: